ARMI Scientists Harness Computing Power to Unlock Biological Networks
A new algorithm developed by ARMI researchers is set to become the gold standard in network analysis. Called “Integrated Value of Influence (IVI),” the algorithm will help systems biology scientists better understand development and disease.
The work, led by PhD student Adrian (Abbas) Salavaty from the Currie Group, was recently published in Patterns– a Cell Press journal dedicated to ground-breaking data science.
“Understanding biology, both healthy and disease states, isn’t just restricted to discoveries made at the lab bench or the clinic bedside. With the technological advances of the past two decades, scientists have been able to harness computing power and the increasing volumes of data we can generate to tease out some fundamental questions in biology. And network science is just one of the methodologies that enable scientists to get deeper insights from the data using a holistic viewpoint,” said Adrian.
Network science is used to investigate complex systems in several fields of research and development, from road traffic to telecommunications to biochemistry. “The way different molecules activate or dampen the expression of other molecules, how they interact with each other- it all forms a network,” explained Adrian. “Disease and disorders arise when such biological networks are altered. So, network science is the perfect tool to help understand biological relationships and propose novel biomarker and drug targets for diseases.”
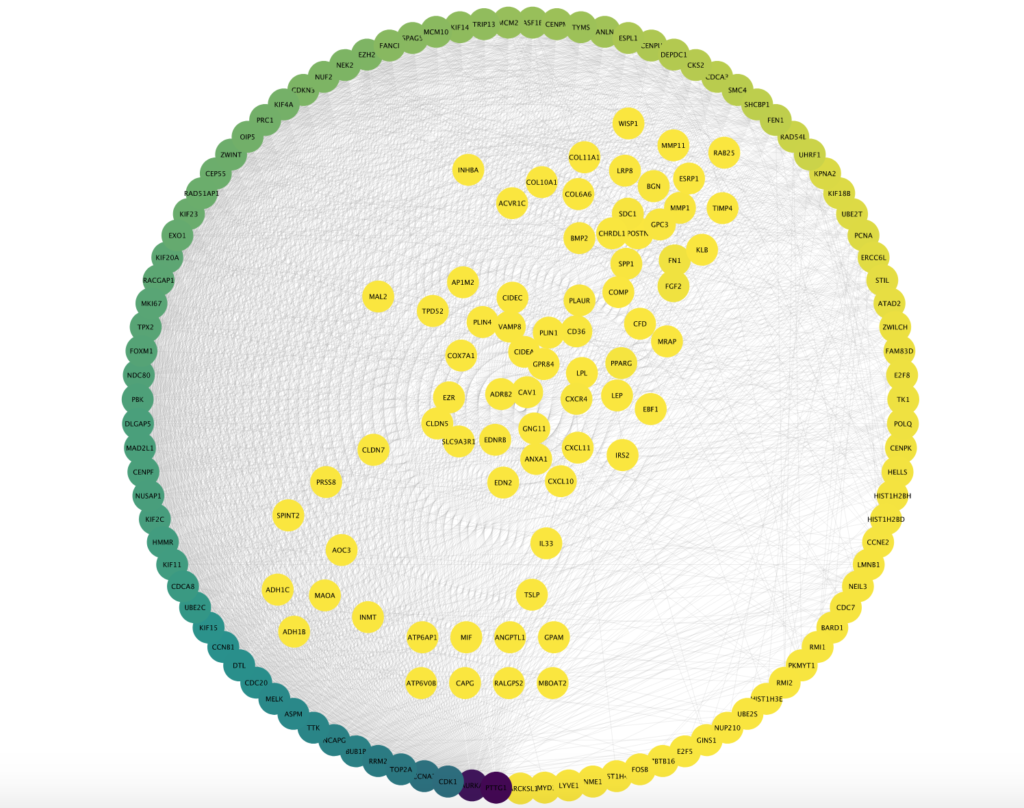
Over the years, algorithms have been developed to identify critical, influential points (also known as nodes) in the network- helping us better understand the critical molecules involved in the development and uncovering potential novel drug targets for treating diseases. Up until now, computational methods have been inadequate in analysing networks that contain a lot of information, called “furballs” in the systems biology world.
To address this major issue, Adrian, alongside supervisors Associate Professor Mirana Ramialison and Professor Peter Currie, worked to create a new and improved algorithm. Adrian undertook a statistical assessment of 200 real-world and simulated networks to develop IVI. When compared against 12 other contemporary influential node identification methods, the IVI algorithm outperformed all other assessed methods.
“Essentially, Adrian has designed a new algorithm to identify the most influential nodes in the network, thereby extracting from the ‘furball’ the information that is crucial for the network,” commented Associate Professor Mirana Ramialison. She continued, “This work has applications beyond systems biology; it could be a great benefit for all future network analyses.”
Congratulations to Adrian, Mirana and Peter!
More information
Click here to read the publication: Salavaty, A., Ramialison, M., & Currie, P. D. Integrated Value of Influence: An Integrative Method for the Identification of the Most Influential Nodes within Networks. Patterns. doi:10.1016/j.patter.2020.100052
The Currie group is curious about the biological mechanisms of the zebrafish, a freshwater fish that is native to South-East Asia. Zebrafish are used in scientific research to understand human genetics and the biological processes of human diseases. For more information on Professor Peter Currie and his group at ARMI, please visit the Currie Group page. You can contact Professor Peter Currie via peter.currie@monash.edu.
The Ramialison Group is a multidisciplinary team of computational and molecular biologists who specialise in genomics. The researchers answer complex questions using new genomic technology and the zebrafish as a model organism. For more information on Associate Professor Mirana Ramialison and her group at ARMI, please visit the Ramialison Group page. You can contact Associate Professor Mirana Ramialison via mirana.ramialison@monash.edu.
