Cracking the Hox Code
The human body navigates movement with astonishing success, displaying locomotor actions that are fluid and coordinated. This ability is the mystery that the McGlinn group is attempting to unravel. Now, they are one step closer with the recent publication of their paper, “A Hox code defines spinocerebellar neuron subtype regionalisation” in Cell Reports.
Corresponding author Associate Professor Edwina McGlinn, ARMI and EMBL Australia group leader noted, “Awareness of relative body position and force required for fluid movement, otherwise known as proprioception, is detected by a complex system of specialised neurons located throughout the body’s muscles, tendons and joints. This system constantly relays information about muscle length, muscle tension and joint position to the spinal cord, information which then needs to be conveyed to the brain for higher order processing. What is lacking in our understanding of proprioception are the molecular underpinnings that shape the formation and function of this neural circuit.”
“This is an exciting step for the field. Our work reveals a pattern of Hox gene expression that gives rise to the molecular, and likely functional, heterogeneity we see in certain types of neurons.” – Dr Victoria Garside
To this end, co-lead authors Drs Eamon Coughlan and Victoria Garside, under the leadership and guidance of Associate Professor McGlinn, have identified a novel role for Hox genes (a family of transcription factors that are fundamental regulators of neural development) in proprioception.
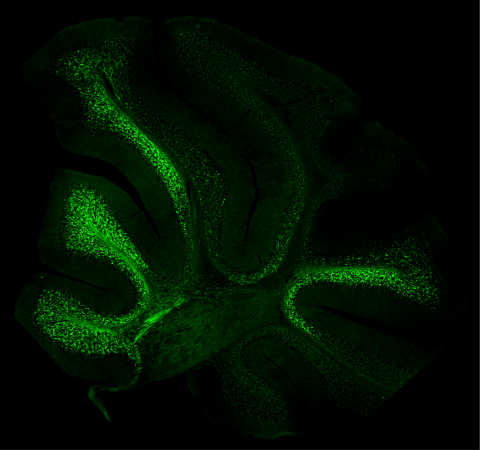
In an elegant and thorough body of work that uses cutting-edge imaging and visualisation techniques and a suite of animal models, this group of scientists and their collaborators from within Australia and in Switzerland have demonstrated extensive and coordinated expression of Hox genes within spinocerebellar neurons, the critical relay neurons that pass proprioceptive information to the brain. The unique “Hox code” they identified within individual spinocerebellar neurons acts as a postcode, telling these cells where along the length of the spinal cord they should reside and how they should function. It was also observed that this expression occurred from embryonic stages through to late adulthood in mice, suggesting a role for Hox genes in the generation and maintenance of spinocerebellar circuits.
“This is an exciting step for the field,” said Dr Garside. “Our work reveals a pattern of Hox gene expression that gives rise to the molecular, and likely functional, heterogeneity we see in certain types of neurons. This work required the development of new tools that enabled us to visualise spinocerebellar neurons with such high resolution, in such granular detail.”
“With these tools, we can now address new questions in the field. How do spinocerebellar neurons navigate their way to the brain? Can we use these tools to track proprioception in disease states?” – Dr Victoria Garside
This project has not only been a finite piece of research in and of itself but provides a solid foundation for further work, many of which take different but exciting directions. “With these tools, we can now address new questions in the field,” says Dr Garside. “How do spinocerebellar neurons navigate their way to the brain? Can we use these tools to track proprioception in disease states?” With proprioception being permanently affected by age and by certain diseases and conditions, such as Parkinson’s disease, it is vital to understand the molecular mediators of movement awareness and explore potential avenues for therapeutic modulation and intervention. Speaking of next steps, Associate Professor McGlinn stated, “We’re excited to follow up and explore the many questions and opportunities this publication has raised.”
Congratulations to Associate Professor McGlinn, Drs Coughlan and Garside and their collaborators on their latest paper!
More information
Click here to read the publication: Coughlan, E. et al. A Hox Code Defines Spinocerebellar Neuron Subtype Regionalization. Cell Reports 29, 2408-2421.e2404, doi:10.1016/j.celrep.2019.10.048 (2019).
The McGlinn Group is focused on elucidating novel gene networks that drive growth and identity in the early embryo. For more information on Associate Professor Edwina McGlinn and the McGlinn Group at ARMI, please visit the McGlinn Group page. You can contact Edwina via edwina.mcglinn@monash.edu.
