Protein conformational diseases such as those leading to neurodegeneration and dementia coincide with the formation of protein aggregates in affected cells/tissues over time leading to premature apoptosis of cells (e.g. brain cells in Alzheimer’s disease and Parkinson’s disease). Often, symptoms linked to these disease conditions become apparent only in the long-lived, aging communities and thus largely limits our ability to develop preventive therapeutics. The Nillegoda group is probing attractive new proteostasis-based directions for future therapeutic interventions that could potentially slow and/or reverse neurodegeneration.
Research
Misfolding/aggregation of cellular proteins is amplified upon exposure to various proteotoxic stresses (e.g. oxidative damage; heat shock), chronic disease conditions and aging. The Nillegoda group utilizes cutting-edge in vitro and in vivo techniques to study chaperone-based protein quality control machineries mediating cell repair. A main focus is currently placed on dissecting the mechanism and regulation of a new class of human protein disaggregases that helps clear aggregated protein junk from cells. These studies will provide valuable insight into the complex pathology of neurodegenerative diseases, and help device strategies to boost aggregate clearance and reduce neurotoxicity in brain cells.
Open positions in the Nillegoda Group
Postdoctoral Fellows
Postdoctoral candidates with a keen interest in proteostasis, cell repair, molecular chaperones and neurodegeneration are invited to directly contact Dr. Nillegoda at nadinath.nillegoda@monash.edu. Applicants should include a cover letter stating research interests, CV and their most recent or pending publications. Prospective postdocs are expected to have a strong publication record.
International and Domestic PhD students
We are currently looking for outstanding PhD students with a strong background in cell biology, biochemistry, biomedical sciences, immunology, neuroscience, or in a closely related field to drive several exciting projects in the lab. Applicants are required to enroll and become graduate research students at Monash University via Monash Graduate Research Program.
International candidates: The successful applicants that secure Monash PhD Scholarships typically have a) MSc with a GPA >3.5 b) 1-2x 1st author publications in Q1 ranked (SJR) journals. Click following link for further information: http://www.armi.org.au/careers-education/higher-degree-research
Domestic candidates: H1 Honours degree (or equivalent) or Masters degree in cell biology, biochemistry, biomedical sciences, immunology, neuroscience, or in a closely related field. Interested students are encouraged to send their CV and academic transcripts to nadinath.nillegoda@monash.edu and arrange a meeting.
For more information: Visit Supervisor Connect at the Faculty of Medicine Nursing and Health Sciences, Monash University.
Honours/Masters Students
We have several positions for undergraduate and MSc students who are part of ARMI and Monash University. Students interested in rotating in the lab please contact Dr. Nillegoda to discuss internships and potential projects. We have several short-term projects that could develop into exciting graduate research work (at PhD level).
Read our guide: The ABCs in choosing a good lab

Funding Acknowledgements
- We are very excited to have received a new NHMRC Investigator Grant (Emerging Leadership) for 2021
- We are grateful to Baillieu and Sarah Myer and the Yulgilbar Foundation for believing in our research on Alzheimer’s disease.


Group Members





Research Themes
- Investigating the underlying molecular and regulatory mechanisms that modulate aggregate clearance in human cells
- Understanding (patho)physiological impact of Hsp70-based protein disaggregation in human health and disease (focus – neurodegenerative diseases)
- Unraveling the interplay between protein aggregation, clearance and immune responses in neurodegenerative diseases
- Development of pharmacological and synthetic biological approaches to control chaperone activity in cells
Protein Repair
The Nillegoda Laboratory is studying chaperone-based cell repair mechanisms with a specific interest in neurodegeneration and aging.
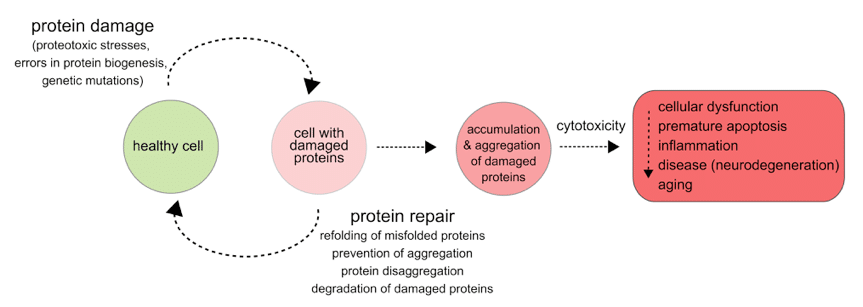
Emerging studies show molecular chaperones that help maintain proteostasis play a critical role in modulating protein conformational diseases in humans ranging from neurodegenerative diseases, metabolic disorders (e.g. Type II diabetes) to cancer. How protein quality control pathways are dysfuncting or misregulated, and understanding what are the molecular and phenotypic consequences of such events is essential for developing effective therapeutic strategies for many of the protein aggregation disorders that have no cures at the moment.
Protein aggregation in neurodegeneration
Progressive neurodegenerative disorders such as Alzheimer’s disease (AD), Parkinson’s disease (PD) and Huntington’s disease (HD) show distinct clinical features, but at molecular level, these conditions are characterized by protein misfolding and aggregation that modulate disease progression and severity by a gain-of-toxicity mechanism(s). Such mechanisms lead to disruption of protein synthesis, trafficking, degradation, and interfere with cell signaling, membrane integrity and could also potentially trigger inflammation. Accumulation of aggregated proteins such as tau and Aβ in AD and related tauopathies, α-synuclein (α-syn) in PD, and polyQ proteins in HD act as hallmarks of these diseases. There is a renewed vigor to develop therapeutics to reduce protein aggregation in the hope of delaying or even halting the progression of these diseases. Towards that end, our lab combines the use of genetics, molecular, cellular, biochemical, and pharmacological methods with animal models of disease to study how protein repair pathways can be boosted to promote the rescue of degenerating or damaged brain cells and prevent their premature apoptosis.
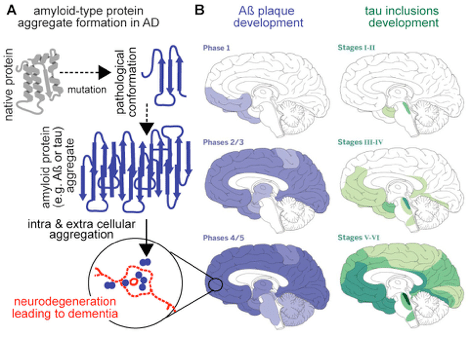
Protein disaggregation
Cellular protein quality control includes all processes that ensure proper protein folding and thus prevent the toxic consequences of protein misfolding and aggregation. Our ground-breaking research uncovered a new class of protein aggregate solubilizing machines (disaggregases) constituted by the Heat shock protein 70 (Hsp70) chaperone system in human cells. This discovery introduced a previously unacknowledged cellular repair activity fundamentally important for counteracting protein aggregation and associated cytotoxicities in disease. A main focus within the group revolves around conceiving novel strategies to activate the human protein disaggregation pathway (PDP). Our work not only generates fundamental knowledge in protein repair mechanisms and stress recovery pathways in human cells, but also an opportunity in translational medicine aimed at clearing toxic aggregated protein junk associated with disease.
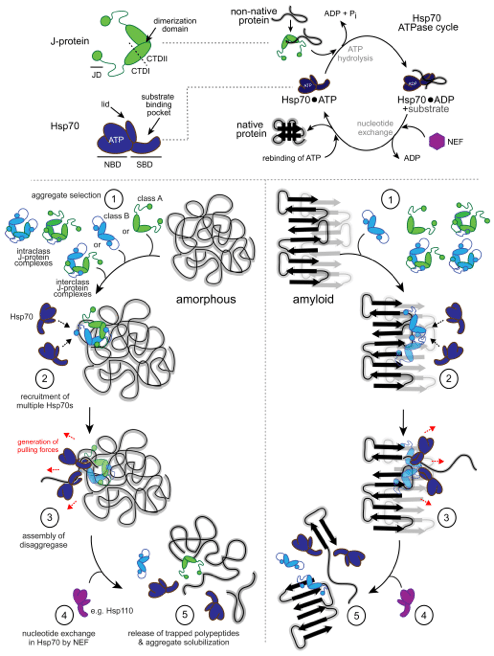
J-domain proteins in proteostasis
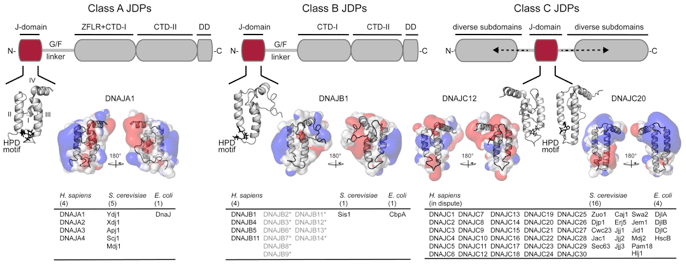
Our lab is particularly interested in the J-domain protein (JDP; J-proteins; Hsp40) family of co-chaperones (classed as A, B, & C according to domain architecture) that drives ATP hydrolysis in Hsp70 and provides substrate specificity. The JDPs form the largest and the most diverse family of co-chaperones in human cells and supports a plethora of Hsp70 dependent and independent proteostasis functions. Though critically important, the lion’s share of the members of this family remains uncharacterized. Very little is known about how these Hsp70 targeting factors bind and select substrates for Hsp70 and help drive distinct cellular processes. The regulation of these co-chaperones could switch “on” or “off” distinct chaperoning activities of Hsp70 that supports transcription/ translation, cell signaling, growth, survival, differentiation, identity, and proliferation; important cellular functions commonly altered in disease. Further, viruses and other pathogens hijack JDPs to support their life cycles in human cells during infection. All in all, these co-chaperones pose as attractive drug targets, especially in light of their emerging association with human health and disease.
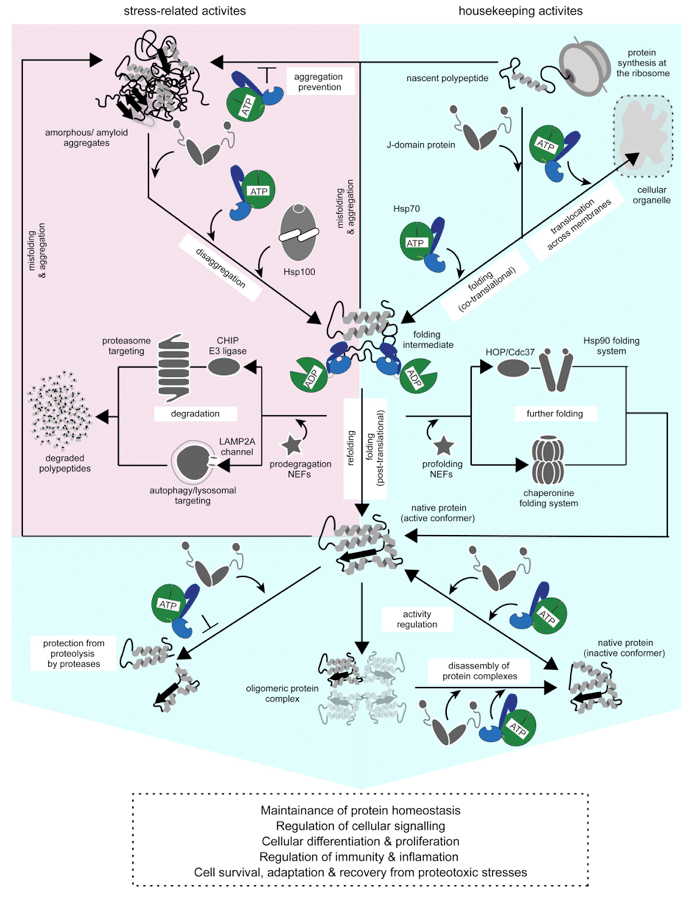
Featured Publications
| Authors | Title | Published In |
|---|---|---|
van Oosten-Hawle P, Backe SJ, Ben-Zvi A, Bourboulia D, Brancaccio M, Brodsky J, Clark M, Colombo G, Cox MB, De Los Rios P, Echtenkamp F, Edkins A, Freeman B, Goloubinoff P, Houry W, Johnson J, LaPointe P, Li W, Mezger V, Neckers L, Nillegoda NB, Prahlad V, Reitzel A, Scherz-Shouval R, Sistonen L, Tsai FTF, Woodford MR, Mollapour M, Truman AW. |
Second Virtual International Symposium on Cellular and Organismal Stress Responses, September 8-9, 2022. |
Cell Stress Chaperones. 2023 Jan 5. doi: 10.1007/s12192-022-01318-5. Online ahead of print. |
Zhang R, Malinverni D, Cyr DM, Rios PL, Nillegoda NB. |
J-domain protein chaperone circuits in proteostasis and disease. |
Trends Cell Biol. 2023 Jan;33(1):30-47. doi: 10.1016/j.tcb.2022.05.004. |
Cox D, Ang CS, Nillegoda NB, Reid GE, Hatters DM. |
Hidden information on protein function in censuses of proteome foldedness. |
Nat Commun. 2022 Apr 14;13(1):1992. doi: 10.1038/s41467-022-29661-2. |
Mathangasinghe Y, Fauvet B, Jane SM, Goloubinoff P, Nillegoda NB. |
The Hsp70 chaperone system: distinct roles in erythrocyte formation and maintenance. |
Haematologica. 2021 Jun 1;106(6):1519-1534. doi: 10.3324/haematol.2019.233056. |
Wentink AS, Nillegoda NB, Feufel J, Ubartaitė G, Schneider CP, De Los Rios P, Hennig J, Barducci A, Bukau B. |
Molecular dissection of amyloid disaggregation by human HSP70. |
Nature. 2020 Nov 11. doi: 10.1038/s41586-020-2904-6. Online ahead of print. |
Serlidaki D, van Waarde MAWH, Rohland L, Wentink AS, Dekker SL, Kamphuis MJ, Boertien JM, Brunsting JF, Nillegoda NB, Bukau B, Mayer MP, Kampinga HH, Bergink S. |
Functional diversity between HSP70 paralogs caused by variable interactions with specific co-chaperones. |
J Biol Chem. 2020 May 22;295(21):7301-7316. doi: 10.1074/jbc.RA119.012449. Epub 2020 Apr 13. PMID: 32284329 |
Faust O, Abayev-Avraham M, Wentink AS, Maurer M, Nillegoda NB, London N, Bukau B, Rosenzweig R. |
HSP40 proteins use class-specific regulation to drive HSP70 functional diversity. |
Nature. 2020 Nov 11. doi: 10.1038/s41586-020-2906-4. Online ahead of print. |
Alberts N, Mathangasinghe Y, Nillegoda NB. |
In Situ Monitoring of Transiently Formed Molecular Chaperone Assemblies in Bacteria, Yeast, and Human Cells. |
J Vis Exp. 2019 Sep 2;(151). doi: 10.3791/60172. |
Rosenzweig R*, Nillegoda NB*, Mayer MP* and Bukau B*. (* Equal contribution) |
The Hsp70 chaperone network. |
Nat Rev Mol Cell Biol. 2019 Jun. doi: 10.1038/s41580-019-0133-3. Epub 2019 Jun 28. |
Nillegoda NB, Wentink AS, Bukau B. |
Protein disaggregation in multicellular organisms. |
Trends Biochem Sci. 2018 Apr;43(4):285-300. doi: 10.1016/j.tibs.2018.02.003. Epub 2018 Feb 28. |
Kirstein J, Arnsburg K, Scior A, Szlachcic A, Guilbride DL, Morimoto RI, Bukau B, Nillegoda NB. |
In vivo properties of the disaggregase function of J-proteins and Hsc70 in Caenorhabditis elegans stress and aging. |
Aging Cell. 2017 Dec;16(6):1414-1424. doi: 10.1111/acel.12686. Epub 2017 Oct 10. |
Nillegoda NB, Stank A, Malinverni D, Alberts N, Szlachcic A, Barducci A, De Los Rios P, Wade RC, Bukau B. |
Evolution of an intricate J-protein network driving protein disaggregation in eukaryotes. |
Elife. 2017 May 15;6. pii: e24560. doi: 10.7554/eLife.24560. |
Nillegoda NB, Kirstein J, Szlachcic A, Berynskyy M, Stank A, Stengel F, Arnsburg K, Gao X, Scior A, Aebersold R, Guilbride DL, Wade RC, Morimoto RI, Mayer MP, Bukau B. |
Crucial HSP70 co-chaperone complex unlocks metazoan protein disaggregation. |
Nature. 2015 Aug 13;524(7564):247-51. doi: 10.1038/nature14884. Epub 2015 Aug 5. |
Gao X, Carroni M, Nussbaum-Krammer C, Mogk A, Nillegoda NB, Szlachcic A, Guilbride DL, Saibil HR, Mayer MP, Bukau B. |
Human Hsp70 disaggregase reverses Parkinson’s-linked α-Synuclein amyloid fibrils. |
Mol Cell. 2015 Sep 3;59(5):781-93. doi: 10.1016/j.molcel.2015.07.012. Epub 2015 Aug 20. |
Nillegoda NB, Bukau B. |
Metazoan Hsp70-based protein disaggregases: emergence and mechanisms. |
Front Mol Biosci. 2015 Oct 9;2:57. doi: 10.3389/fmolb.2015.00057. eCollection 2015. |
Rampelt H, Kirstein-Miles J, Nillegoda NB, Chi K, Scholz SR, Morimoto R, Bukaua B. |
Metazoan Hsp70 machines use Hsp110 to power protein disaggregation. |
EMBO J. 2012;31(21):4221-4235. doi: 10.1038/emboj.2012.264. |
More Publications
| Authors | Title | Published In |
|---|---|---|
Kampinga HH, Andreasson C, Barducci A, Cheetham ME, Cyr D, Emanuelsson C, Genevaux P, Gestwicki JE, Goloubinoff P, Huerta-Cepas J, Kirstein J, Liberek K, Mayer MP, Nagata K, Nillegoda NB, Pulido P, Ramos C, De Los Rios P, Rospert S, Rosenzweig R, Sahi C, Taipale M, Tomiczek B, Ushioda R, Young JC, Zimmermann R, Zylicz A, Zylicz M, Craig EA, Marszalek J. |
Function, evolution, and structure of J-domain proteins. |
Cell Stress and Chaperones. 2018 Nov 26. doi: 10.1007/s12192-018-0948-4. Epub ahead of print 2018 Nov 26. |
Substrate binding by the yeast Hsp110 nucleotide exchange factor and molecular chaperone Sse1 is not obligate for its biological activities. |
Mol Biol Cell. 2017 Jul 15;28(15):2066-2075. doi: 10.1091/mbc.E17-01-0070. Epub 2017 May 24. |
|
Żwirowski S, Kłosowska A, Obuchowski I, Nillegoda NB, Piróg A, Ziętkiewicz S, Bukau B, Mogk A, Liberek K. |
Hsp70 displaces small heat shock proteins from aggregates to initiate protein refolding. |
EMBO J. 2017 Mar 15;36(6):783-796. doi: 10.15252/embj.201593378. Epub 2017 Feb 20. |
Hsieh TY, Nillegoda NB, Tyedmers J, Bukau B, Mogk A, Kramer G. |
Monitoring protein misfolding by site-specific labeling of proteins in vivo. |
PLoS One. 2014 Jun 10;9(6):e99395. doi: 10.1371/journal.pone.0099395. eCollection 2014. |
A network of ubiquitin ligases is important for the dynamics of misfolded protein aggregates in yeast. |
J Biol Chem. 2012 Jul 6;287(28):23911-22. doi: 10.1074/jbc.M112.341164. Epub 2012 May 16. |
|
Mandal AK, Theodoraki MA, Nillegoda NB, Caplan AJ. |
Role of molecular chaperones in biogenesis of the protein kinome. |
Methods Mol Biol. 2011;787:75-81. doi: 10.1007/978-1-61779-295-3_6. |
Nillegoda NB, Theodoraki MA, Mandal AK, Mayo KJ, Ren HY, Sultana R, Wu K, Johnson J, Cyr DM, Caplan AJ. |
Ubr1 and Ubr2 function in a quality control pathway for degradation of unfolded cytosolic proteins. |
Mol Biol Cell. 2010 Jul 1;21(13):2102-16. doi: 10.1091/mbc.e10-02-0098. Epub 2010 May 12. |
Mandal AK, Gibney PA, Nillegoda NB, Theodoraki MA, Caplan AJ, Morano KA. |
Hsp110 chaperones control client fate determination in the hsp70-Hsp90 chaperone system. |
Mol Biol Cell. 2010 May 1;21(9):1439-48. doi: 10.1091/mbc.e09-09-0779. Epub 2010 Mar 17. |
Mandal AK, Nillegoda NB, Chen JA, Caplan AJ. |
Ydj1 protects nascent protein kinases from degradation and controls the rate of their maturation. |
Mol Cell Biol. 2008 Jul;28(13):4434-44. doi: 10.1128/MCB.00543-08. Epub 2008 Apr 28. |
Jones J, Wu K, Yang Y, Guerrero C, Nillegoda NB, Pan ZQ, Huang L. |
A targeted proteomic analysis of the ubiquitin-like modifier nedd8 and associated proteins. |
J Proteome Res. 2008 Mar;7(3):1274-87. doi: 10.1021/pr700749v. Epub 2008 Feb 5. |
Mandal AK, Lee P, Chen JA, Nillegoda NB, Heller A, DiStasio S, Oen H, Victor J, Nair DM, Brodsky JL, Caplan AJ. |
Cdc37 has distinct roles in protein kinase quality control that protect nascent chains from degradation and promote posttranslational maturation. |
J Cell Biol. 2007 Jan 29;176(3):319-28. doi: 10.1083/jcb.200604106. Epub 2007 Jan 22. |



